PIBAS FedSPARQL - user guide
PIBAS FedSPARQL allows users to easily run predefined Federated SPARQL queries across multiple data sources (Bio2RDF, Chem2Bio2RDF, EMBL-EBI and CPCTAS). As an advanced feature, system can detect similar data items (instance matching).
Basic steps:
- Select topic
- Select subtopic
- Select template
- Enter keyword
- Run query
| Example of running template Find targets for the drug |
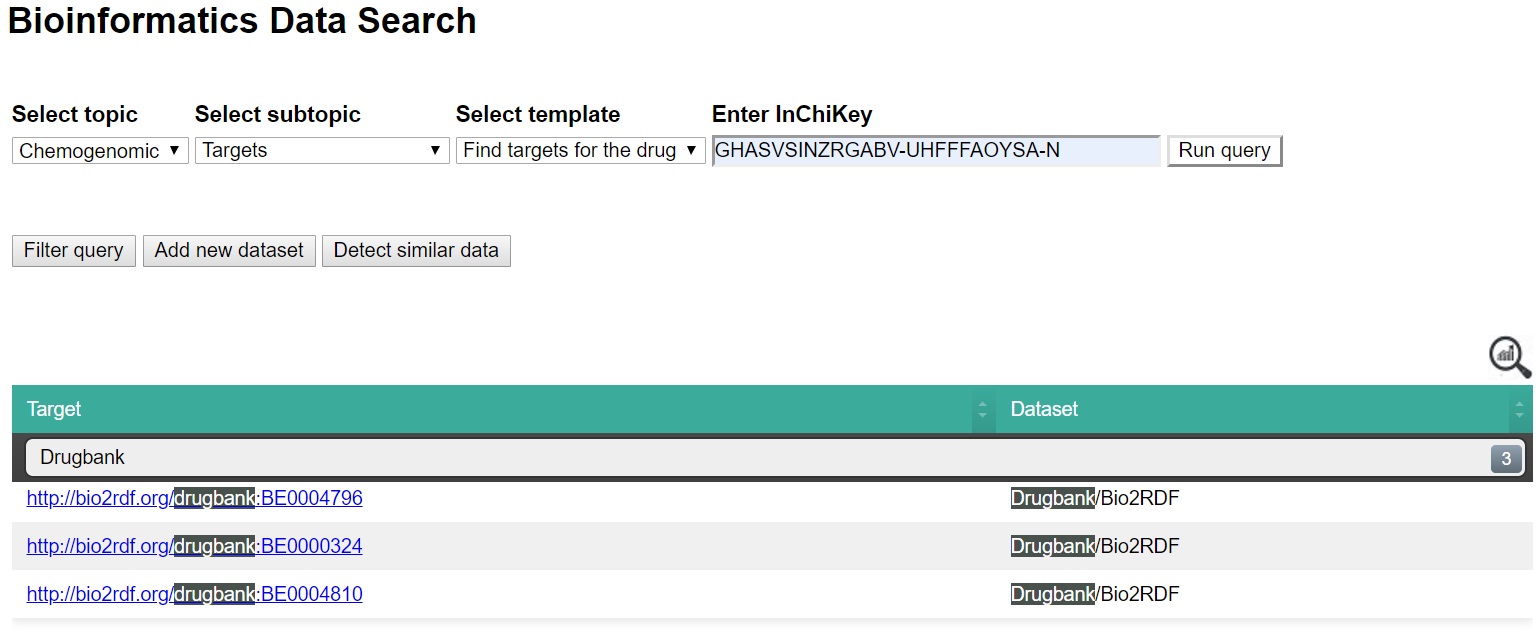
|
Additional features
1. Add new dataset: Users need to enter information such as: dataset name, initiative name, dataset link, comment, endpoint URL and pattern query. Requirement pattern query should match to a selected topic and template. Conversance with SPARQL is necessary for this step, because the added dataset may be unfamiliar to bioinformatics community. Following our use case, pattern query must contain variable target that fits to the name of running template. Pattern query variable is visible in the right corner of the pop-up window for adding of new dataset. Users have an option to maker their data public.
| Example of adding test dataset |
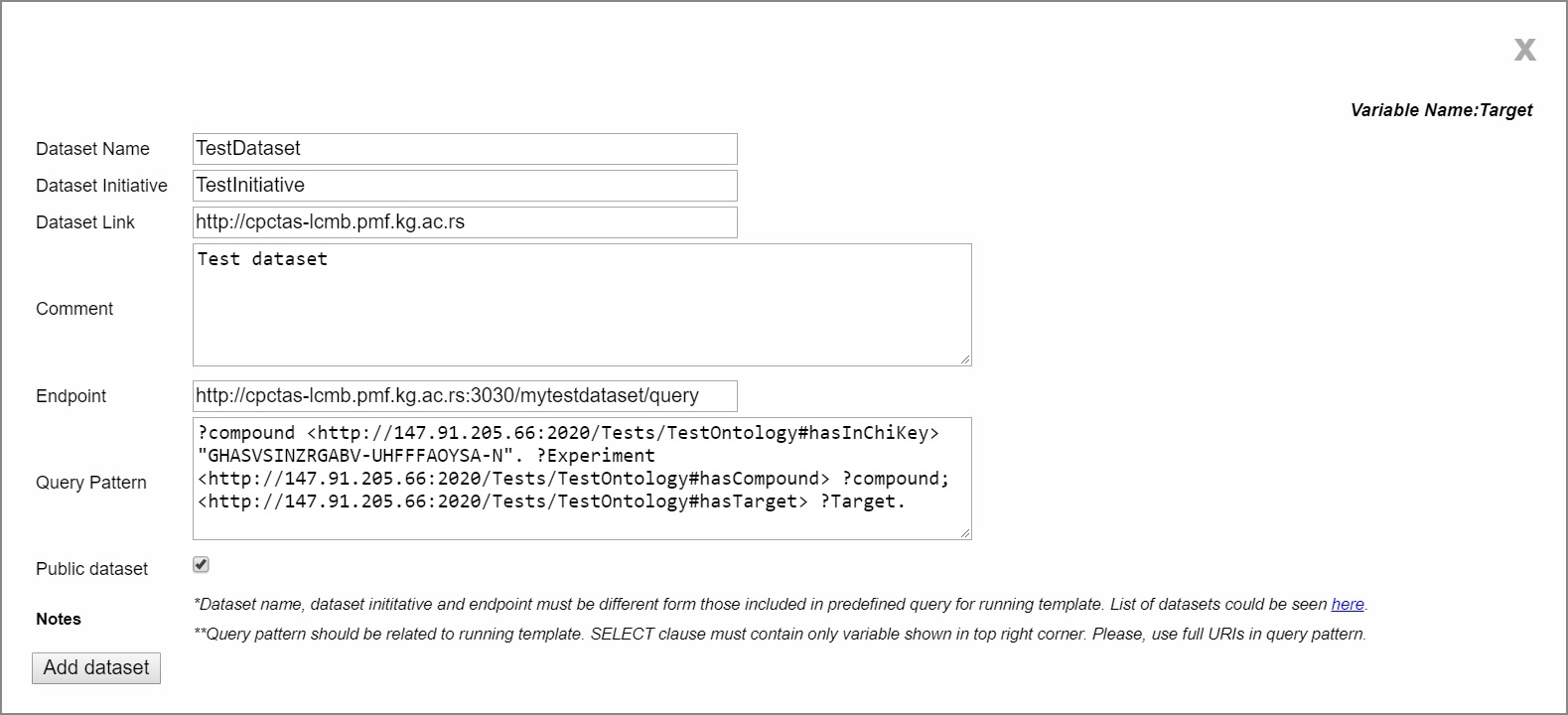 |
| Result after adding newdataset |
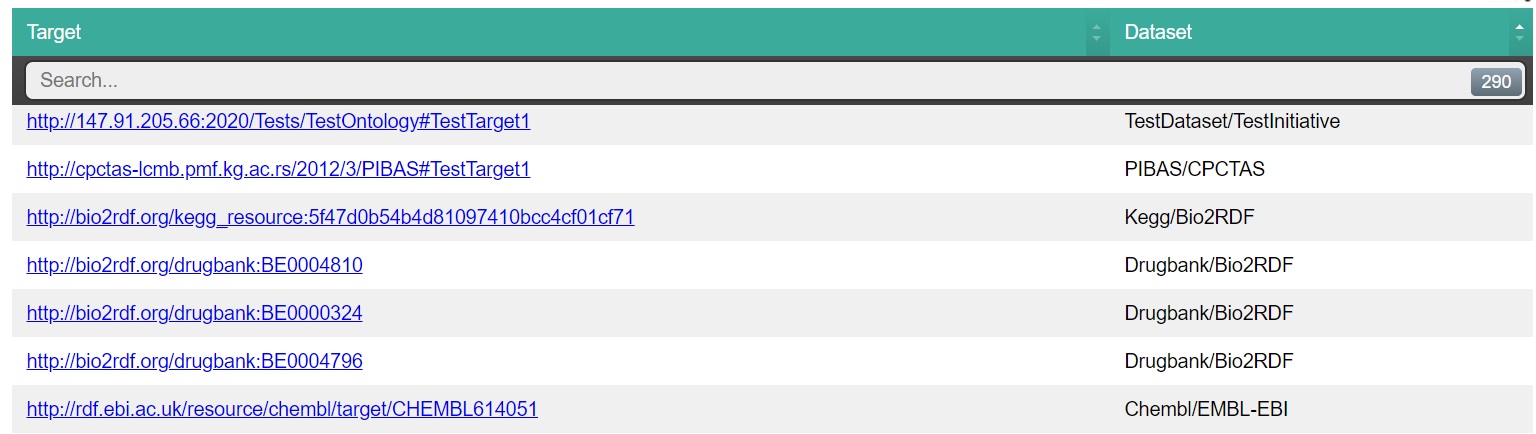 |
2. Dynamic query filtering: This feature allows users to improve their queries using the underlying structure of datasets without the prior knowledge of their structure. Each dataset used in query is assigned to accordion element. Accordion elements are labeled with dataset name and initiative name. The names are linked, so the end-user can directly explore the given dataset or initiative through their websites or public endpoints. By clicking on accordion element, it gets expanded and automatically populated with the list of properties, which are dependent on selected template and topic. This list is generated by running background dynamic SPARQL query. Each property, listed in accordion element, has a hyperlink to the web page with its description. Analyzing properties, users can decide whether some of them are relevant for obtaining additional information. Each property can be added to the query, by selecting it, and then, the query button "Run query" changes to "Run new query".
| Accordion elements for dynamic query filtering |
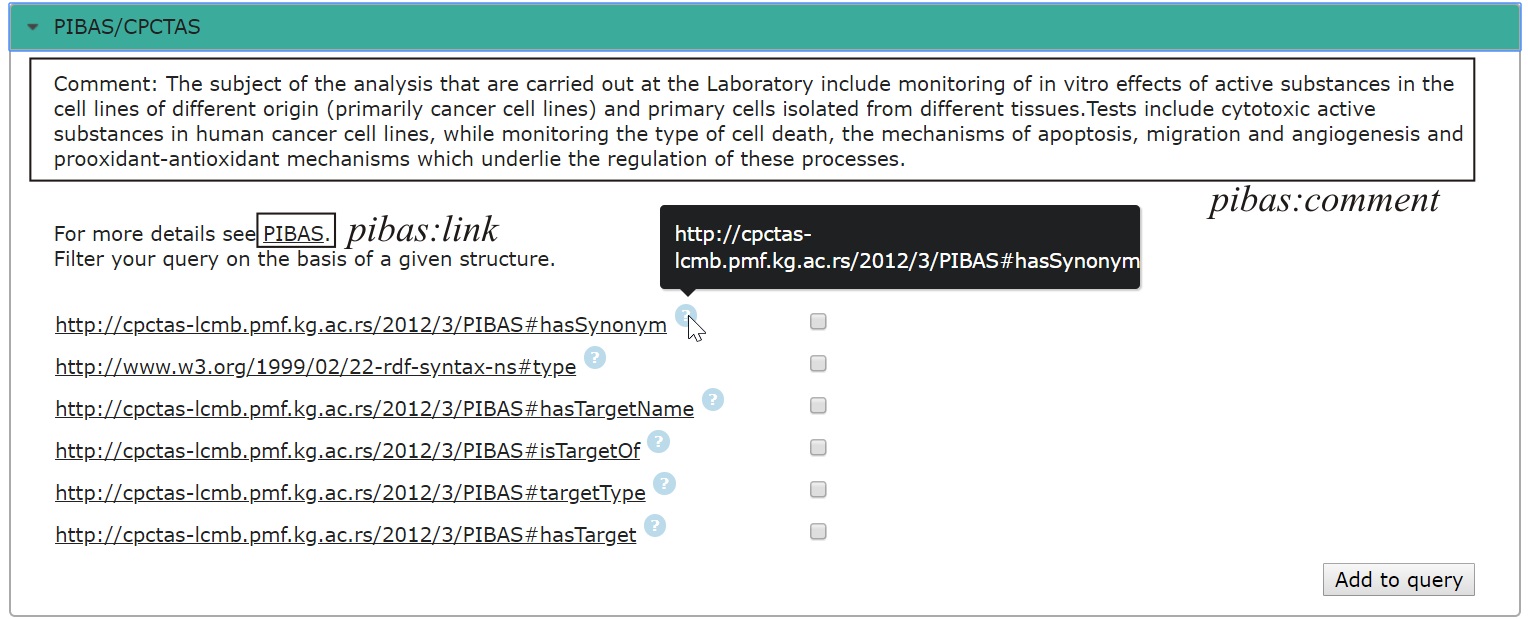 |
| A sample result table after dynamic query filter |
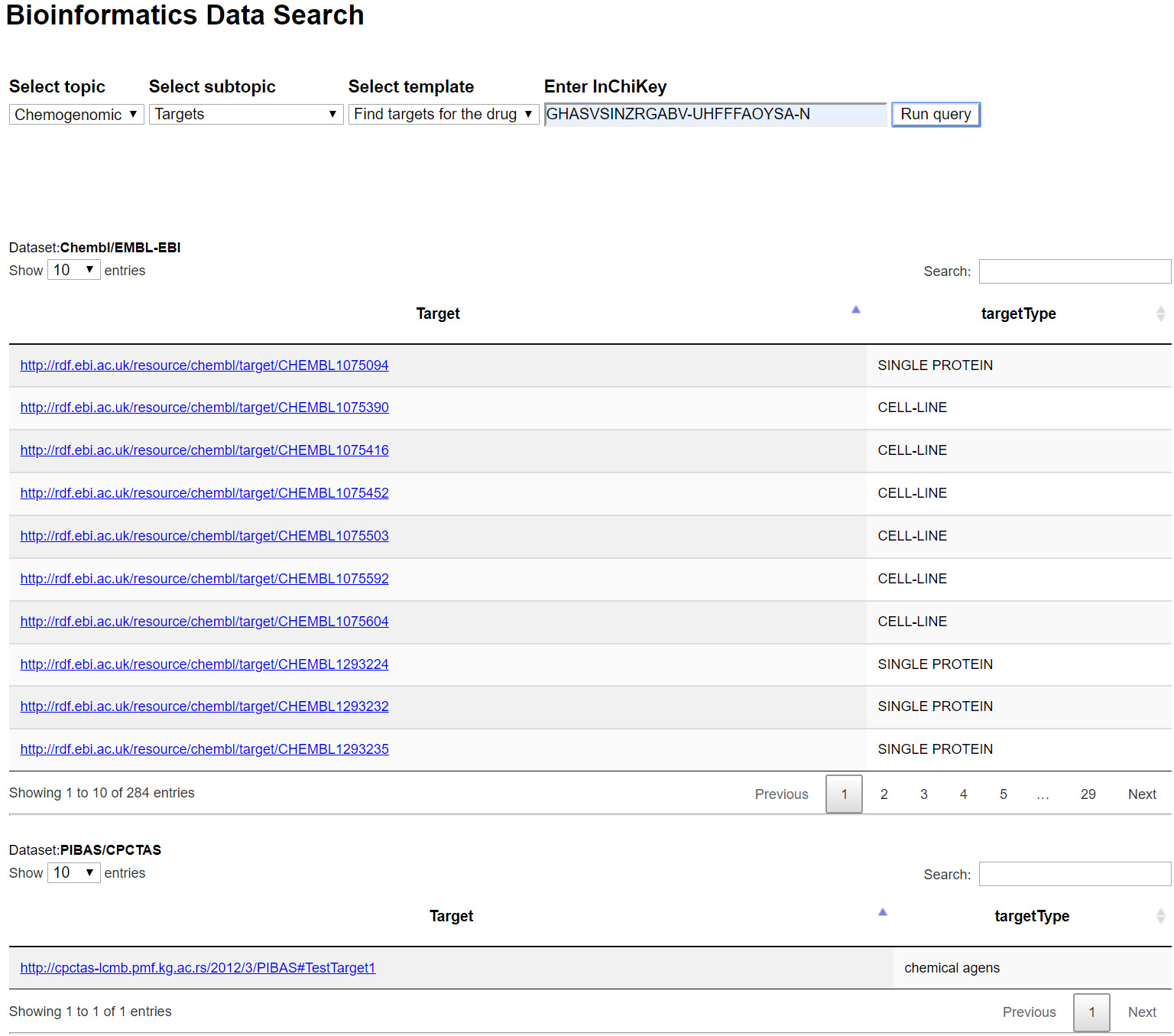 |
3. Detection of similar data items: This option can be applied on the results of predefined queries as well as on the results retrieved after adding a new dataset. This feature could be disabled for some templates in DataSources ontology. It is important for some Biology and Chemogenomic topics. Researchers can find the most similar targets by click on button "Detect similar data items".
| Similar data items after adding of new dataset |
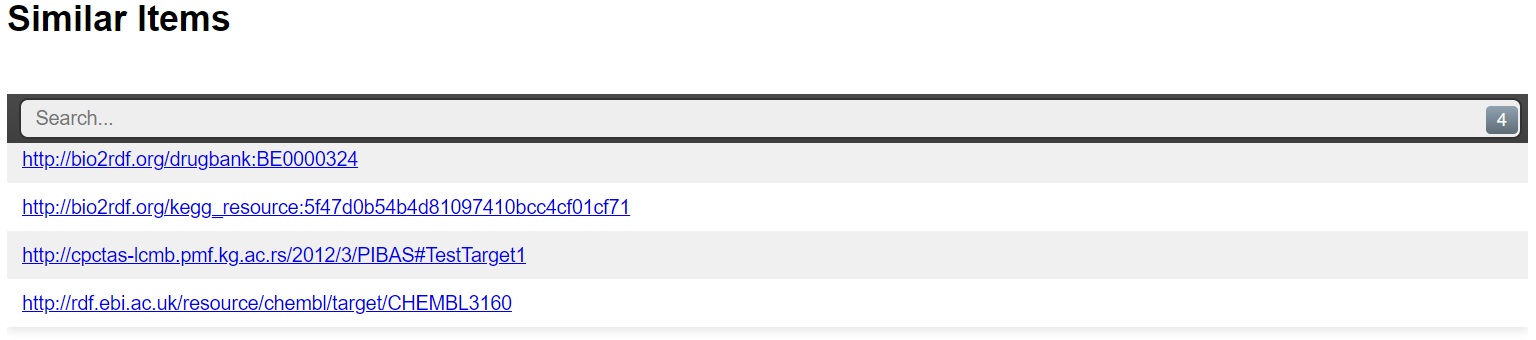
|
You can watch PIBASFed SPARQL demo here.